Assignment 5
1. First, to prepared the group of sequence which want to do phylogenetic tree. In this case I selected 11 sequence for do this test.
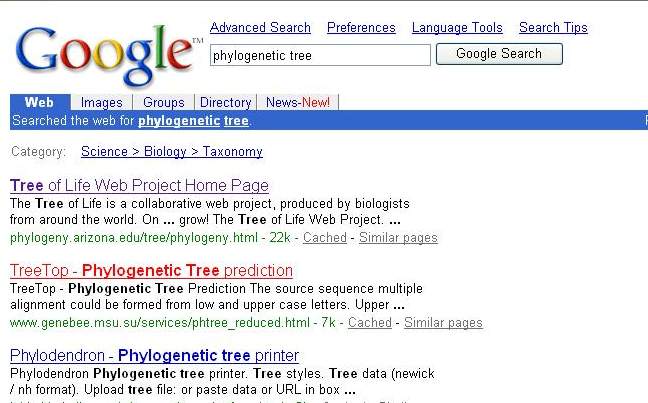
2. Then I search the tool which can make the phylogenetic tree from Google web by the key word "phylogenetic tree".
3. Select the tool that is interest. The tree top web (http://www.genebee.msu.su/services/phtree_reduce.html) will open.

4. Copy all sequence that we want to make phylogenetic
tree from notepad data.
5. Paste all sequence to correct box and
change some parameter for the appropriation. Then click submit and
wait for the result.
6. The result will show in the folowing pictures.Save
this data for analyze furthur.
7.Then I will change the nucleotide
sequences to the amino acid sequences, so I will search this tool in google
web by use key word"protein translate".
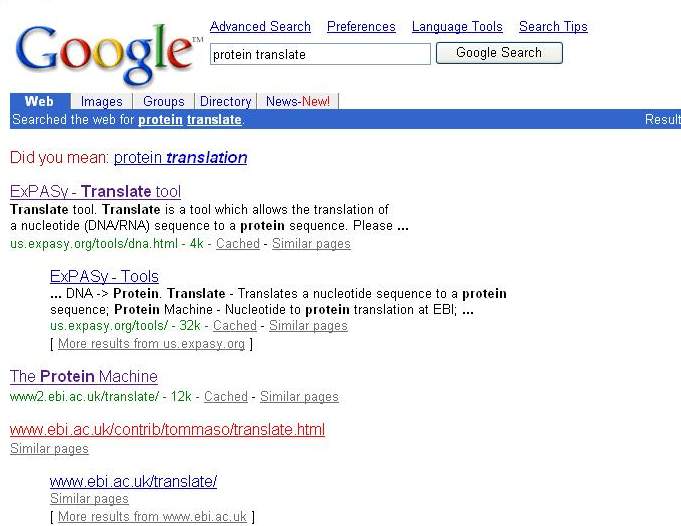
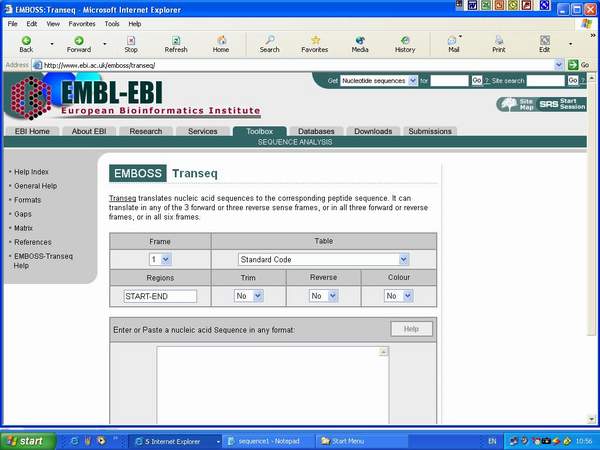
8. Select the web that
is interest. The EMBOSS Transeq web(http://www.ebi.ac.uk/emboss/transeq/)
will open.
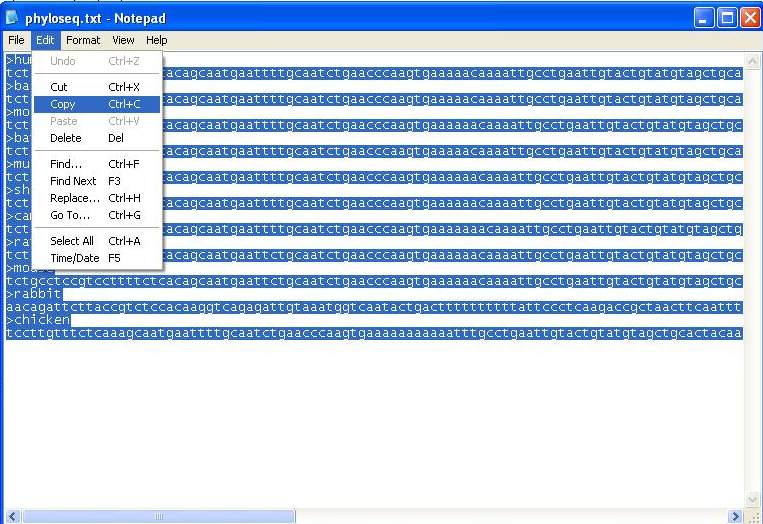
9. Copy the nucleotide sequences from notepad data.
10. Paste all sequence to correct box and
change some parameter for the appropriation. Then click RUN and wait
for the result.
11. The result will show in the following picture.
12. Copy this result the amino acid sequences to notepad and save for use furthur.
13. Open the Tree top phylogenetic tree web and copy this amino acid sequences.
14. Paste that sequence to the correct box, click submit and wait for the result.
15. The result will show in the following pictures.
Conclution
When compare the result of phylogenetic tree from the
nucleotide sequences to amino acid sequences,both are similarly result.
In my opinion,the nucleotide sequence is easier to analyze than the amino acid sequence because it didn't change to amino acid sequence before do the phylogenetic tree or it has less step than other.
In addition, I think, because of many triplet
nucleotide sequence can translate to same amino acid.So the nucleotide sequence
should have more value to tell the relationship between different organisms,
animals or species than by use amino acid sequence.