
![]() Assignment
3:
Assignment
3:
Deadline: Dec31,2002 at 16.30
Please notify insturtor once
you complete this assigment uisng this e-mail address: assign3@learnbioinfo.org
***************************************************************
Download the same gene sequence
from five species/strains of bacteria available in NCBI-Entrez web site
- Build up a multiple alignment
- Design a common primer pair and one probe for five sequences, which
all templates should be amplified and probed by this probe and primer
set.
![]()
I interest in pennicillin resistant bacteria and I want to focus on Streptococcus pneumoniae and other bacteria that can resist to pennicillin. Then I so steps by step be low.
1) Search for interested penicillin resistant baterias by key word "penicillin resistant" from nucleotides search of NCBI. Look for Streptococcus pneumoniae and click to get sequence from linked page. And scroll down to select other interested 4 strains of bacterias.
2)Copy the sequence to past and save in microsoft word doccument. Keep as back up file for multiple alignment step.

3) Find multiple alignment tool from PIR web page. Put in sequences that kept from step3), FASTA format, all 5 sequences are placed separate by " >". For example, >seq1 atatatatatatatatatatatatctctctcctctctct.............. >seq2 ctcgctsgtatagtgtacgtagctagcta.......>seq3 atatatctctgctgtgatcgatcgt......... , then click "submit" to get result.
4) The result of multiple alignment show below, "tree view" of relation and detail of alignment show more when scroll down.
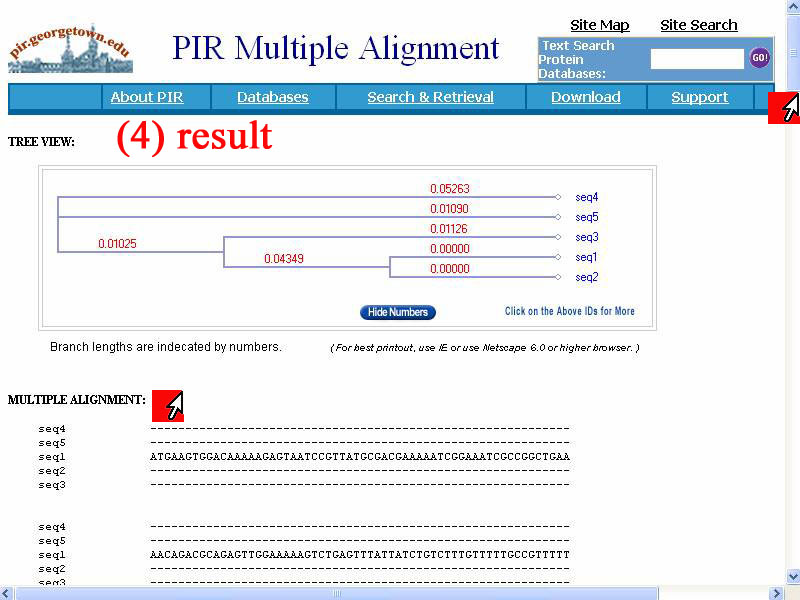
5) The detail of alignment of 5 sequences show as stars (*). Interestingly, consensus sequence of 5 sequences not stared at first of alls but shift in side several hundred bases and very similar base sequence are in midle of all bacterial sequences.
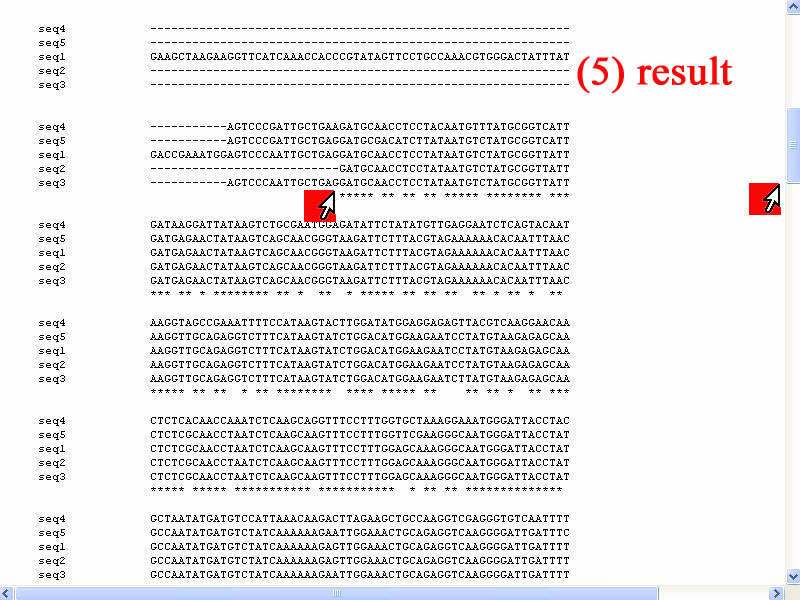
6) In the midle of sequences show continously stars that mean very identical. That good for design common primers and probes specific for all 5 sequence. From this result show very identical as base number 900-1600.

7)For design primers and probe are avilable at "Primer3" web page. From step 6) I know consensus sequence of all bacteria (*) that means all bacterias has very same nucleotides sequence at that specific aria. Pick sequence of seq2 (Streptococcus pneumoniae strain IC100 ) as voluteer instead of all 5 sequences. Select only nucleotide sequene number 961-1051 to put in the box. Check in boxes of "pick" for left and right primers and also probe. In this step you can click "pick primers" immediately after input sequence in order to use default condition or scroll down to specific the condition incase you already have some appropriate condition to use.
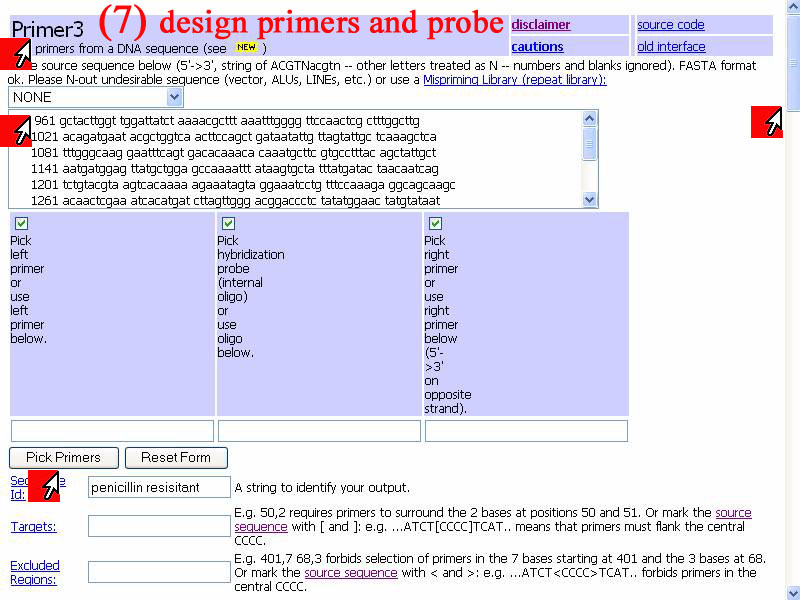
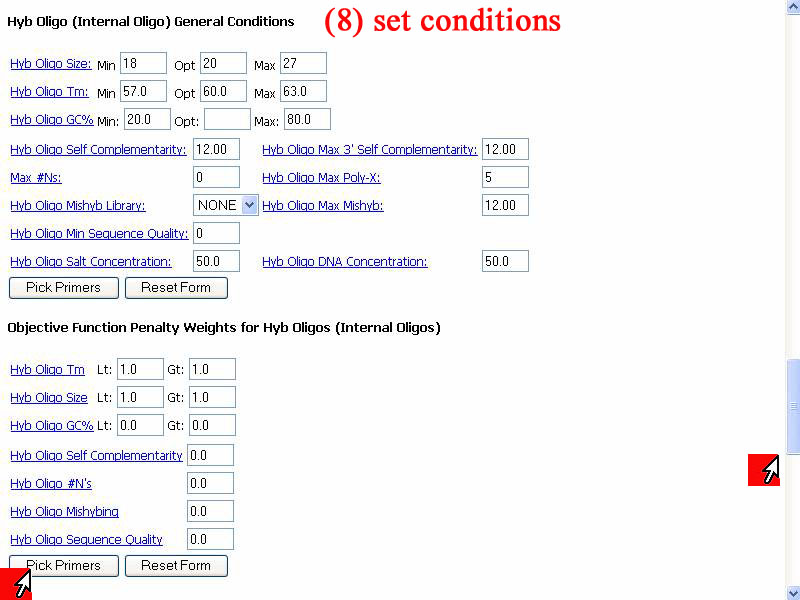
8) To specific conditions of reaction can be done show in window below. After satisfy condition set, click "Pick primers" to get result. In this demonstrate I should default first, if result not satisfy I'll come back to this step and set other appropriate conditions.
9) Output result show in window below. Left and right primers and also 1 primer are designed by "Primer3 tool". The detail of each available and also site of complementary binding of left primer show as ">", right primer shown as "<" and primer shown as "^".

CONCLUSION !!!Select 5 bacterias strains by NCBI nucleotides databases, 3 Streptococcus pneumoniae with different strains and Streptococcus oralis and Streptococcus mitis ,all of it has pennicllin resistant gene that can eventually resist to pennicillin treatment. Do multiple alignment with PIR3 multiple alignment found that consensus sequence of all 5 bacterial sequence are very identical in midle of sequence (nucleotides 961-1051). Then select that sequence from only one bacterial strain for design common primers and probe. From Primer3 design left and riht common primer and 1 primer specific for 5 selected bacterial strains. PCR product of this appropriate set condition is 160 nucleoties that advantage for more specificity but may need more number of cycle time when perform PCR. Designed primer and probe has well characteristic by easy bioinformatic process, by the way if this result is not satisfy it can be easy redesign again by same bioinformatic tools demonstrated above.


