You have genome
sequence (~10,000 bp) of new bacterial specie by shortgun sequence.
Each contigs was cloned to plasmid. Finally, every contigs were sequenced
by automated DNA sequencer.
Suppose, you want to know that this genome sequence contain the qin like
gene which we found in Pseudomonas aeruginosa. What would you do? What
bioinformatics tools that you will use?
![]()
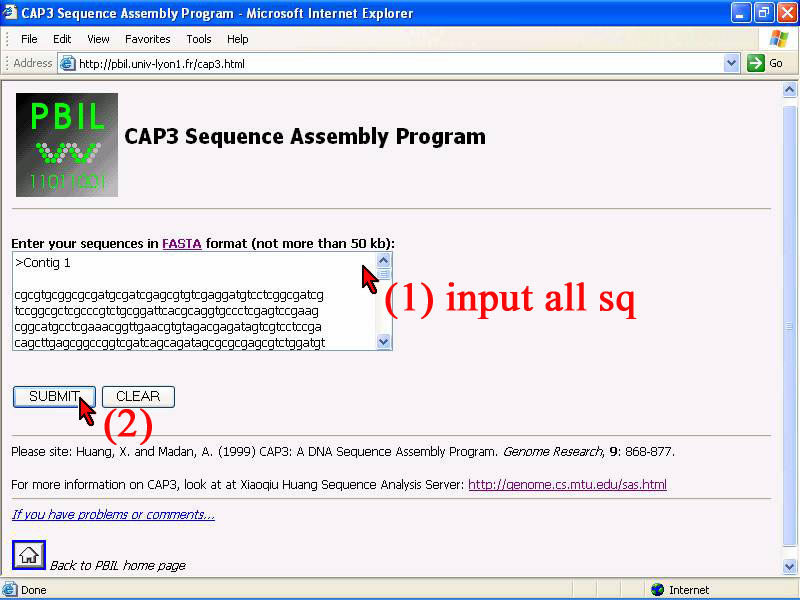
The 10 contiqs given were sequenced splitly from DNA sequencing process that need to be assembled in to 1 full leght DNA sequence by using assembly tool. Find web-base tool as CAP3 tool from internet, it can be easy used for assemble DNA sequences in FASTA format. The process are given by steps below.
1) Do copy all 10 contiqs in FASTA format from source and put in sequence box below.Between contiq separate by ">" and the whole sequence must not biger than 50 Kb.
2) Click "submit" to get result. (,or click "clear" if need to recorrect something)
3) The rusult'll show as summary in result page. Two main importance are "Contiq" and "Assembly deails" as red arrow point. then click to get more reult.
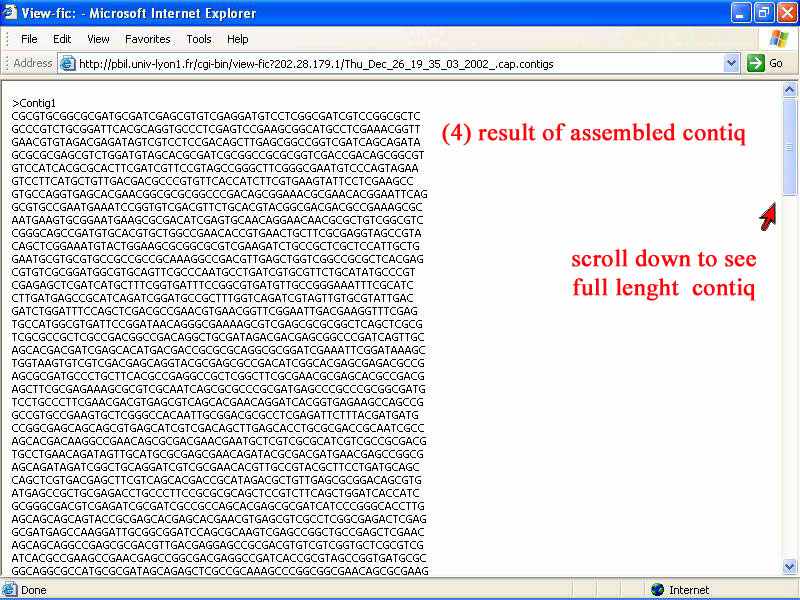
4) The whole new assembled contiq show in this window, available for copy to do allignment for qin gene. In assembly, an overlap sequence was cut off and then continoues sequence be joined together, to give only one new assembled conig shown below.
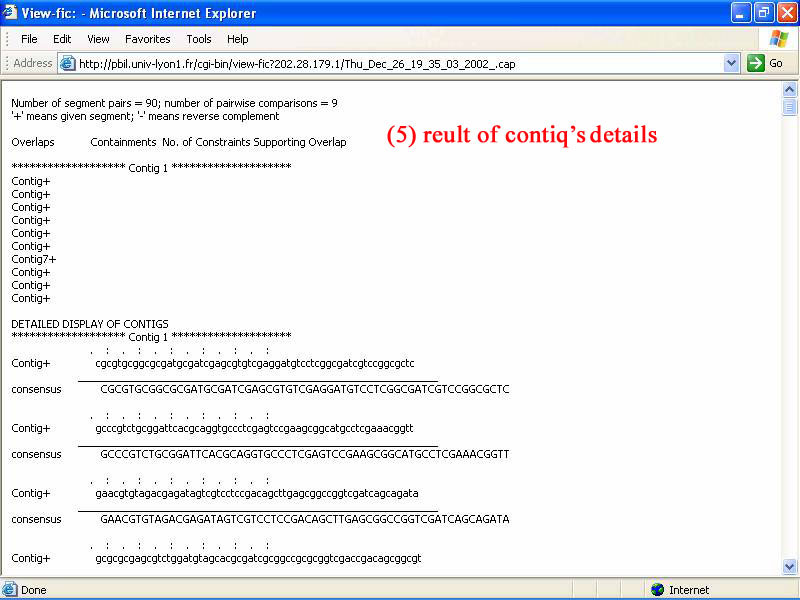
5) The other detail of new contiq available in this window. It can be used for check or find correction of assembly process.
![]() NOW
!! steps of finding for
qine gene in the new contiq be shown sinply by 2 sequences allignment.
The tool that I select for finding qine gene is "Blast
2 sequences" available in Blast
family of NCBI sourch. Open NCBI then select to open Blast 2 sequences
window and do by steps below.
NOW
!! steps of finding for
qine gene in the new contiq be shown sinply by 2 sequences allignment.
The tool that I select for finding qine gene is "Blast
2 sequences" available in Blast
family of NCBI sourch. Open NCBI then select to open Blast 2 sequences
window and do by steps below.
1)Opend form of Blast 2 sequences. Set "Program" as blstn for nucleotide-nucleotide allignment and for other box set as defualt.
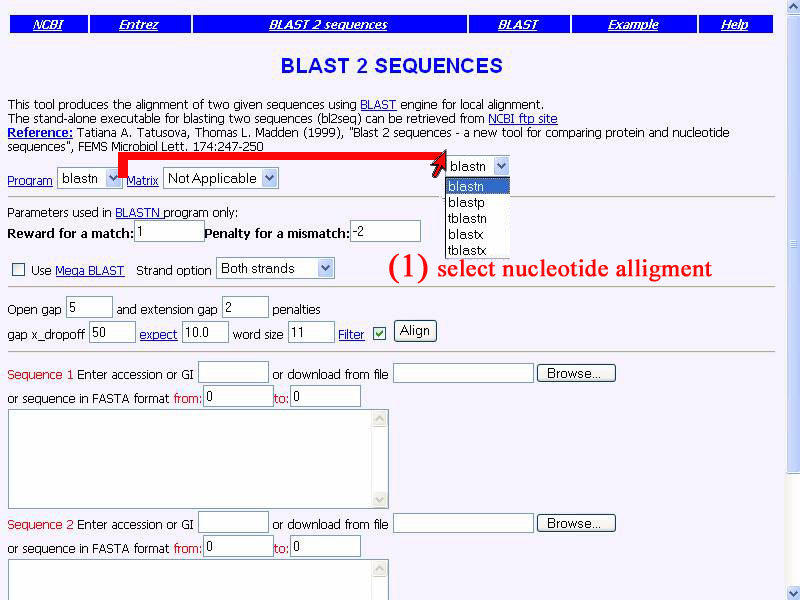
2) Do copy qine gene sequence of P. aeruginosa and put in "sequence 1" box, also for our well-assembled sequence put in "sequence 2 " box.
3) Click "Align".
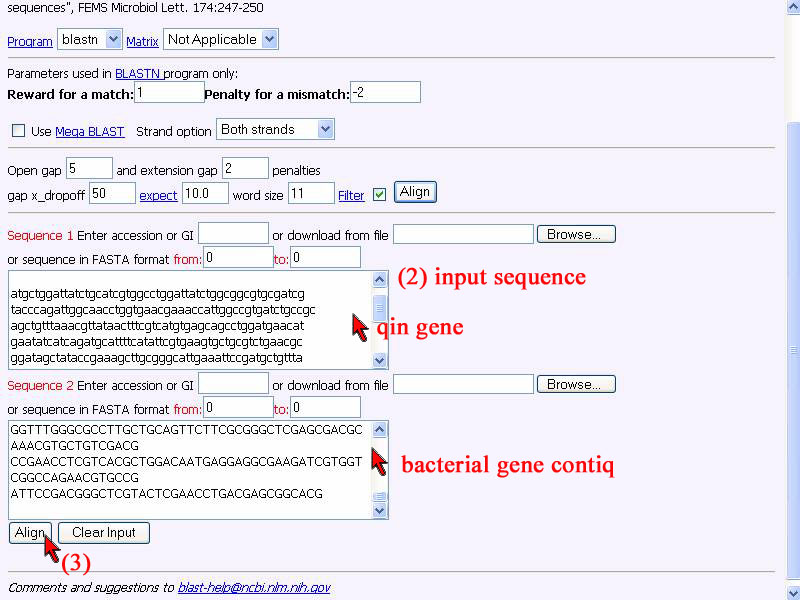
4) Result shown that the identity score of 2 sequence alignment is 83%. Sequence are not 100% identity and many nucleotides not the same, represent thsre is no qin gene in this given bacterial DNA.
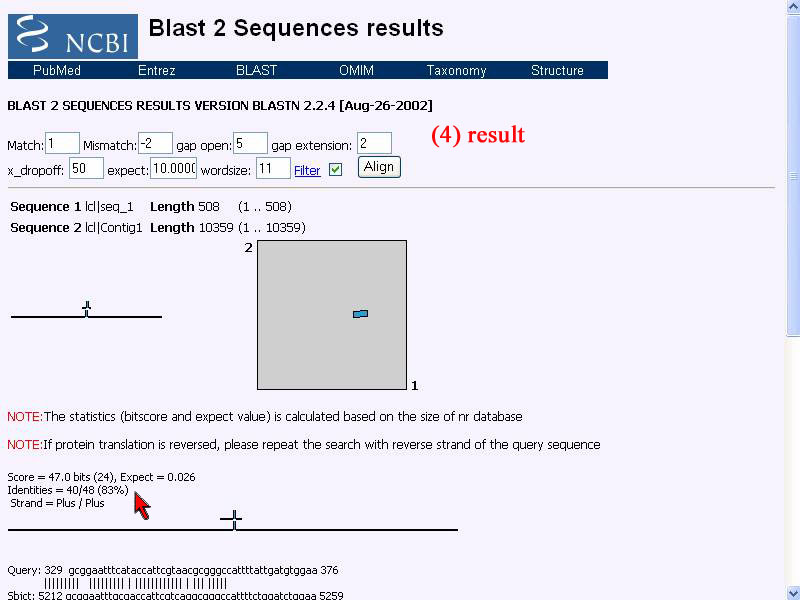

5) However I use "Blastn" to do align that given sequence with nucleotides database and the result show highest score of identity as other gen of B. pseudomelei .
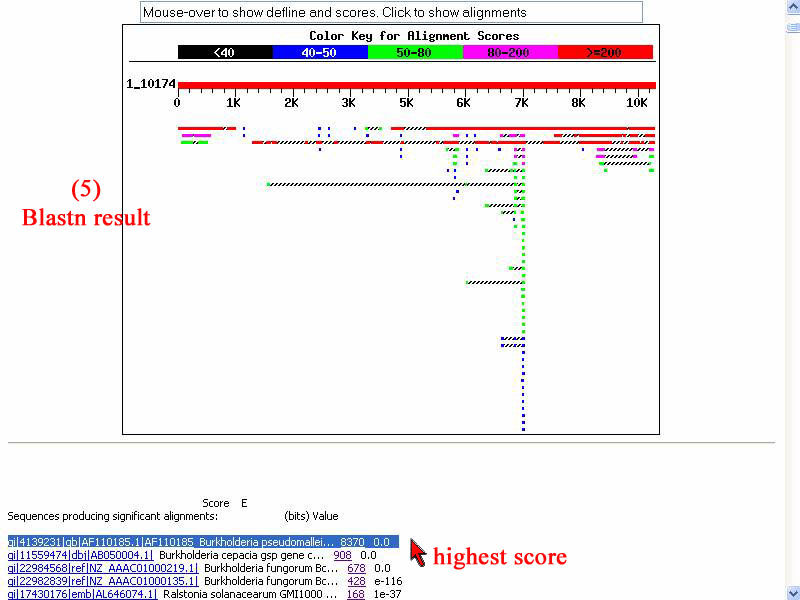
Conclusion : The given bacterial DNA does not have qin gene of P. aeruginosa accoring to result of pairwise alignment and nucleotide alignment with P. aeruginosa qin gene and DNA database respectively.